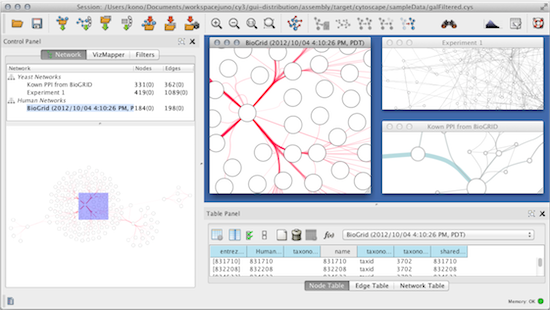 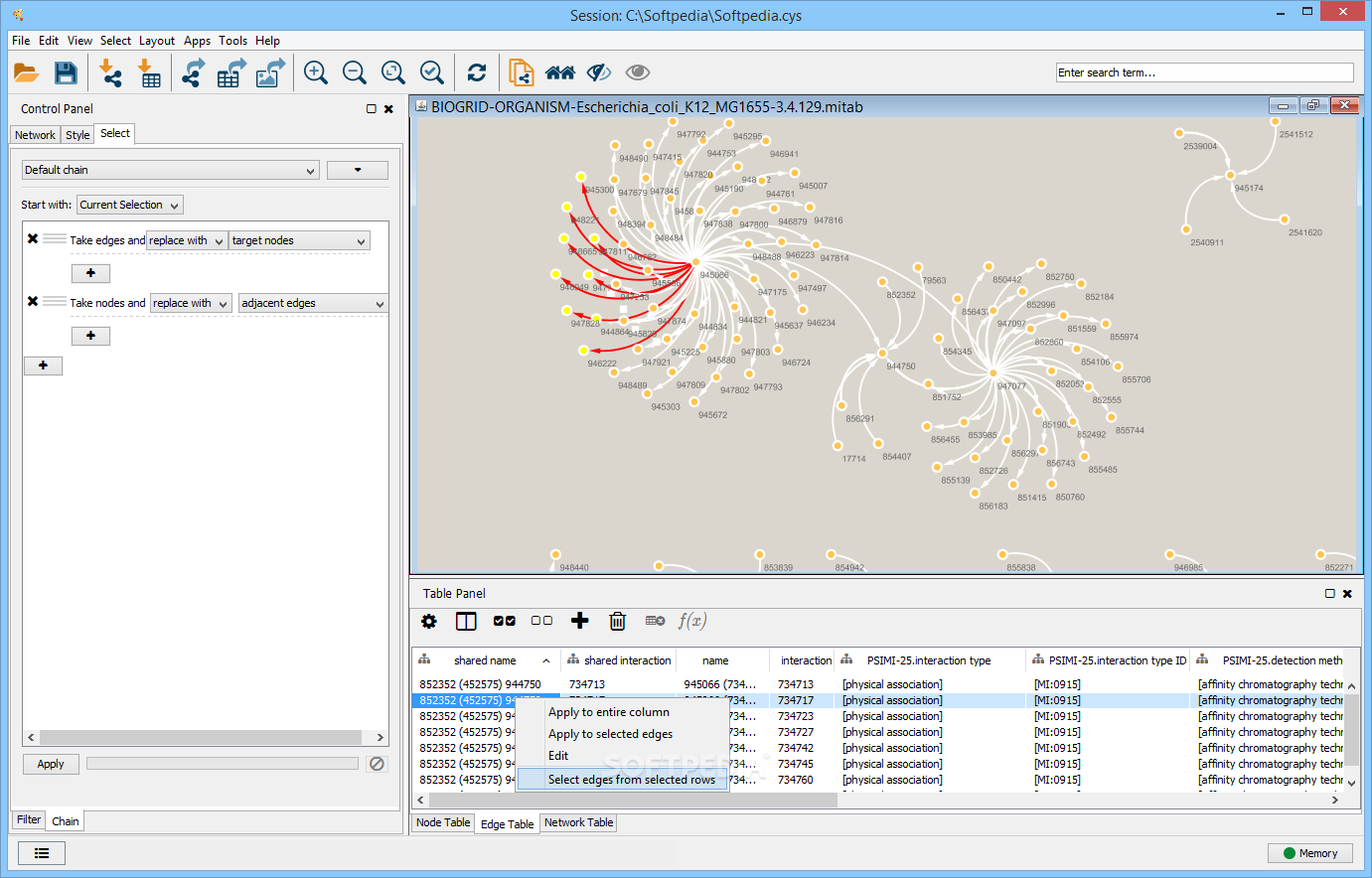
The results showed four potential biomarkers with the highest value in each parameter of topological analysis such as RPA1, USP7, UBC and TERF2, but only RPA1 included in protein module with the highest score of 4.526, while UBC and TERF2 involved in protein module with lower score. Furthermore, biomarkers' candidate expression was confirmed by microarray data from NCBI and analyzed by non-paired t-test. Biomarker was identified by topological analysis, modularity analysis and functional analysis using Cytoscape 3.2.1. This research aims to identify new potential biomarker and those contributions in NPC signalling pathway. Several methods are available for diagnosis but it only indicates the viral titre. Title: Identification of potential biomarkers in nasopharyngeal carcinoma based on protein interaction analysisĪuthors: Yulanda Antonius Didik Huswo Utomo WidodoĪddresses: Faculty of Mathematics and Natural Sciences, Biology Department, Brawijaya University, Malang 65145, Indonesia ' Faculty of Mathematics and Natural Sciences, Biology Department, Brawijaya University, Malang 65145, Indonesia ' Faculty of Mathematics and Natural Sciences, Biology Department, Brawijaya University, Malang 65145, IndonesiaĪbstract: Nasopharyngeal carcinoma (NPC) is malignant tumour that strongly related to Epstein-Barr virus infection. International Journal of Bioinformatics Research and Applications.Inderscience Publishers - linking academia, business and industry through research This study might help to understand the NPC mechanism and develop an appropriate treatment. Moreover, RPA1 has high expression in NPC samples (p < 0.05 FC = 1.07) and mainly related to cell cycle pathway. This study provides a theoretical basis for the follow-up development of medicines for the treatment of HCC.Article: Identification of potential biomarkers in nasopharyngeal carcinoma based on protein interaction analysis Journal: International Journal of Bioinformatics Research and Applications (IJBRA) 2017 Vol.13 No.4 pp.376 - 388 Abstract: Nasopharyngeal carcinoma (NPC) is malignant tumour that strongly related to Epstein-Barr virus infection. Conclusion :Network pharmacology intuitively shows that Radix Paeoniae Rubra has multiple components ,multiple targets and multiple pathways in the treatment of HCC. A total of 561 biological processes ,85 cellular components and 130 molecular functions were obtained by GO annotation. There were 193 nodes and 722 edges in the protein- protein interaction network ,and 95 nodes ,439 edges and 21 core targets were obtained by topological index screening. Twenty-three common targets from both active ingredient targets and HCC disease targets were obtained. 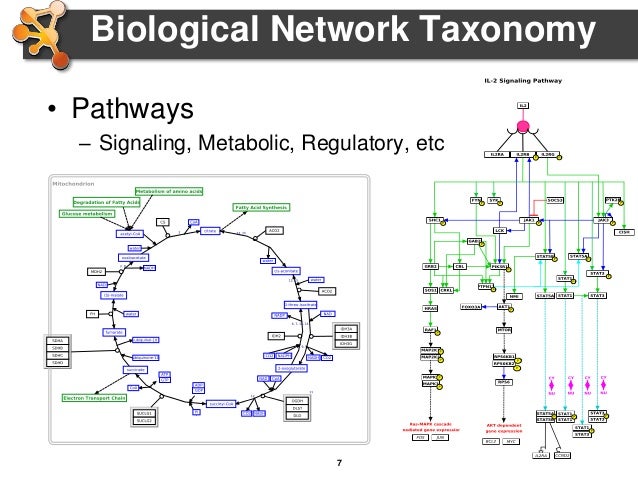
A total of 148 HCC-related disease targets were retrieved from OMIM, DrugBank and TTD databases. Results :A total of 28 compounds and 442 targets from Radix Paeoniae Rubra were obtained from TCMSP database. The mechanism of Radix Paeoniae Rubra on HCC was further analyzed. Protein-protein interaction (PPI) network was constructed by Cytoscape 3.2.1 software, and the core targets were obtained by topological analysis. The String platform was used to collect liver cancer disease-related target proteins, and GO analysis was performed with David database. Then, the common targets from both active ingredient targets and HCC disease targets were obtained. Through OMIM database (http ://),drugbank database (https :// )and TTD database (http ://db./ttd/ ),the corresponding targets of HCC were collected. Methods :Through TCMSP database (https :///tcmspsearch.php ),the chemical constituents of Radix Paeoniae Rubra were searched ,and the targets of Radix Paeoniae Rubra were predicted by STD database and SEA database. Objective :To predict the mechanism of Radix Paeoniae Rubra in the treatment of hepatocellular carcinoma (HCC )by network pharmacology.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
